Inbreeding
Inbreeding occurs when an individual has one or more common ancestors.
The term 'common ancestor' means an ancestor that is present at least once on both sides of an individual's pedigree. It does not mean an ancestor that just 'occurs a lot'.
When an individual has common ancestors, this raises the possibility that for any particular gene locus, both alleles present are in fact physical copies of the exact same allele from one of its common ancestors. The inbreeding coefficient is the value of this probability. Or, put another way, it is the probability that both alleles at any given gene locus are identical by descent.
The inbreeding coefficient as in common use today was first defined in 1922 in a paper by Dr Sewall Wright (Wright, S. "Coefficients of inbreeding and relationship. American Naturalist 56: 330-338, 1922"). For this reason it is often known as 'Wrights Inbreeding Coefficient'.
When an individual has no common ancestors (within the ancestry being considered), its inbreeding coeffficient is zero. The symbol 'F' is commonly used in scientific literature to mean an individual's inbreeding coefficient.
The inbreeding coefficient can be computed by hand only for relatively simple pedigrees. For deep pedigrees it can only realistically be done by computer. Calculating inbreeding for very large pedigrees is a computationally intensive process.
Breeders Assistant can compute the inbreeding coefficient and associated information such as loss of heterozygosity and the relationship coefficient. It can also produce a number of reports of related information, such as common ancestor listings. Much of this information can be easily included in forms such as pedigrees.
Inbreeding options within Breeders Assistant include:
- Configurable number of generations of ancestors to include in the calculation (max 60).
- Configurable precision (number of decimal places reported).
- Whether to include the inbreeding coefficient on pedigrees and trial mating pedigrees (and also for each ancestor if you want).
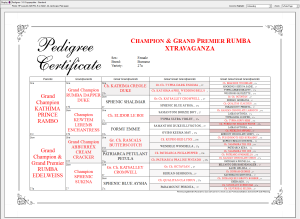
- Whether to include the relationship coefficient on trial mating pedigrees and mating certificates1.
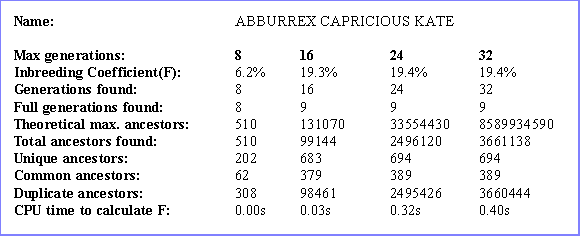
- Generation of related information including: (i) the depth at which the furthest ancestor was found; (ii) how many generations of ancestors were full; (iii) total number of ancestors found; (iv) number of unique ancestors; (v) number of ancestors common to both sides of the pedigree.
Video
Many of the inbreeding features of Breeders Assistant are demonstrated in this video ![]()
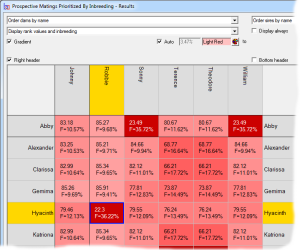
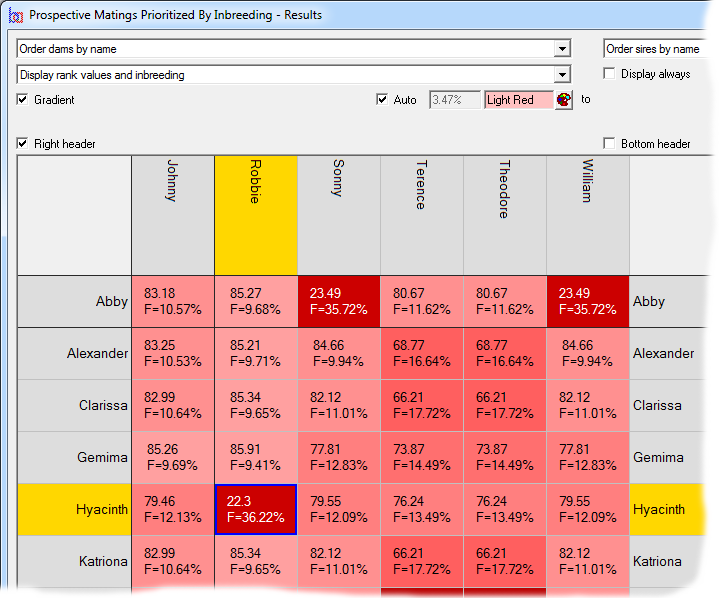
Minimizing Inbreeding
With the Extended Edition it is possible to rank large numbers of prospective matings according to the inbreeding that would result in the offspring. This is an efficient way to select a mating specifically to minimize inbreeding. See video ![]()
It is possible to take additional factors when minimizing prospective mating inbreeding. E.g. if you are a breeder of pedigree animals you may want to select a mating with the deliberate intention of maximizing the influence that a given individual (or individuals) has on the resultant offspring. Or, you may want to discourage matings where specific individuals share a high kinship with the offspring (to discourage traits you wish to breed away from). All this, and more, is available with the of advanced population genetics of the Extended Edition.
Bulk Inbreeding Calculations
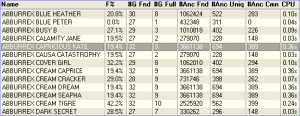
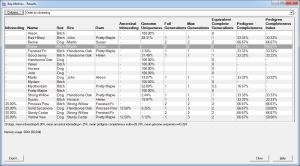
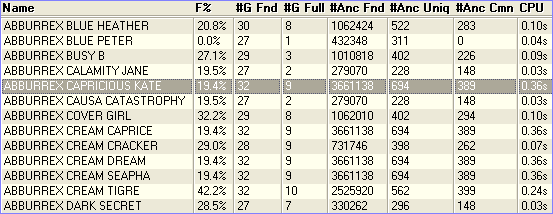
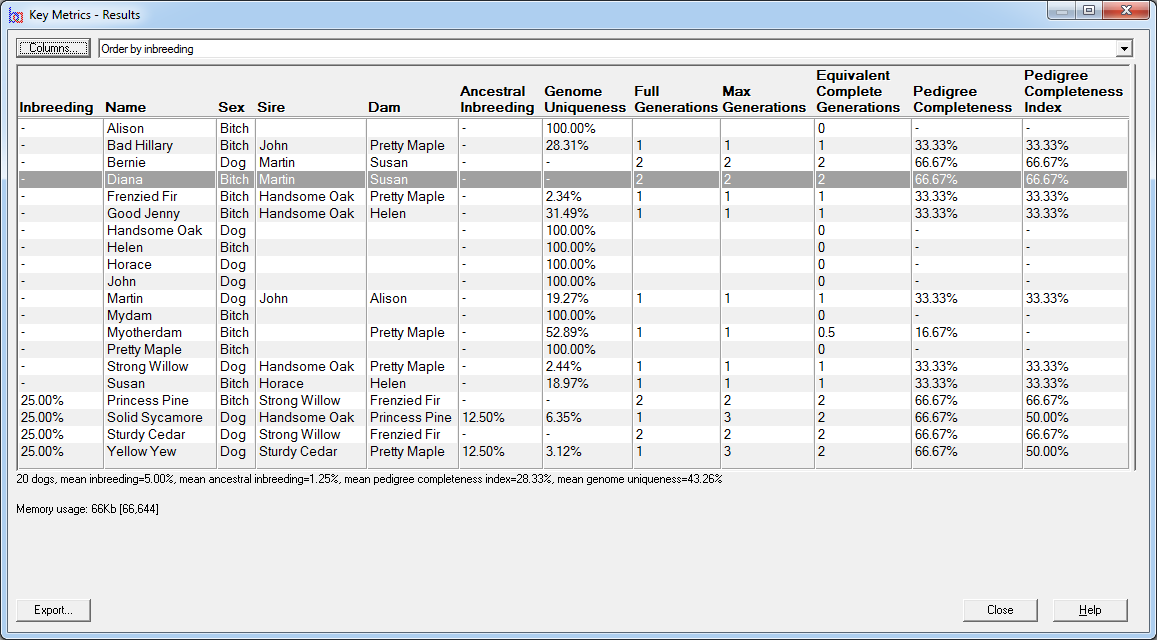
It is possible to tabulate all the inbreeding calculation details given above directly in the main window, for all animals in the database. Just select Inbreeding Table from the View pulldown (just above the animal table). Breeders Assistant rapidly calculates the inbreeding coefficients for the entire database.
Inbreeding coefficients can be exported easily using text/CSV export. Simply choose Database|Export and use the Export Layouts button to create an export layout that includes the inbreeding coefficient in addition to other columns (such as name, date of birth etc). Then export the records you need to a text file using the chosen layout.
Tagging and Highlighting
It is possible to tag the common ancestors of an animal, or multiple animals, to a chosen number of generations. Use the Tag|Advanced|Tag Ancestors option.
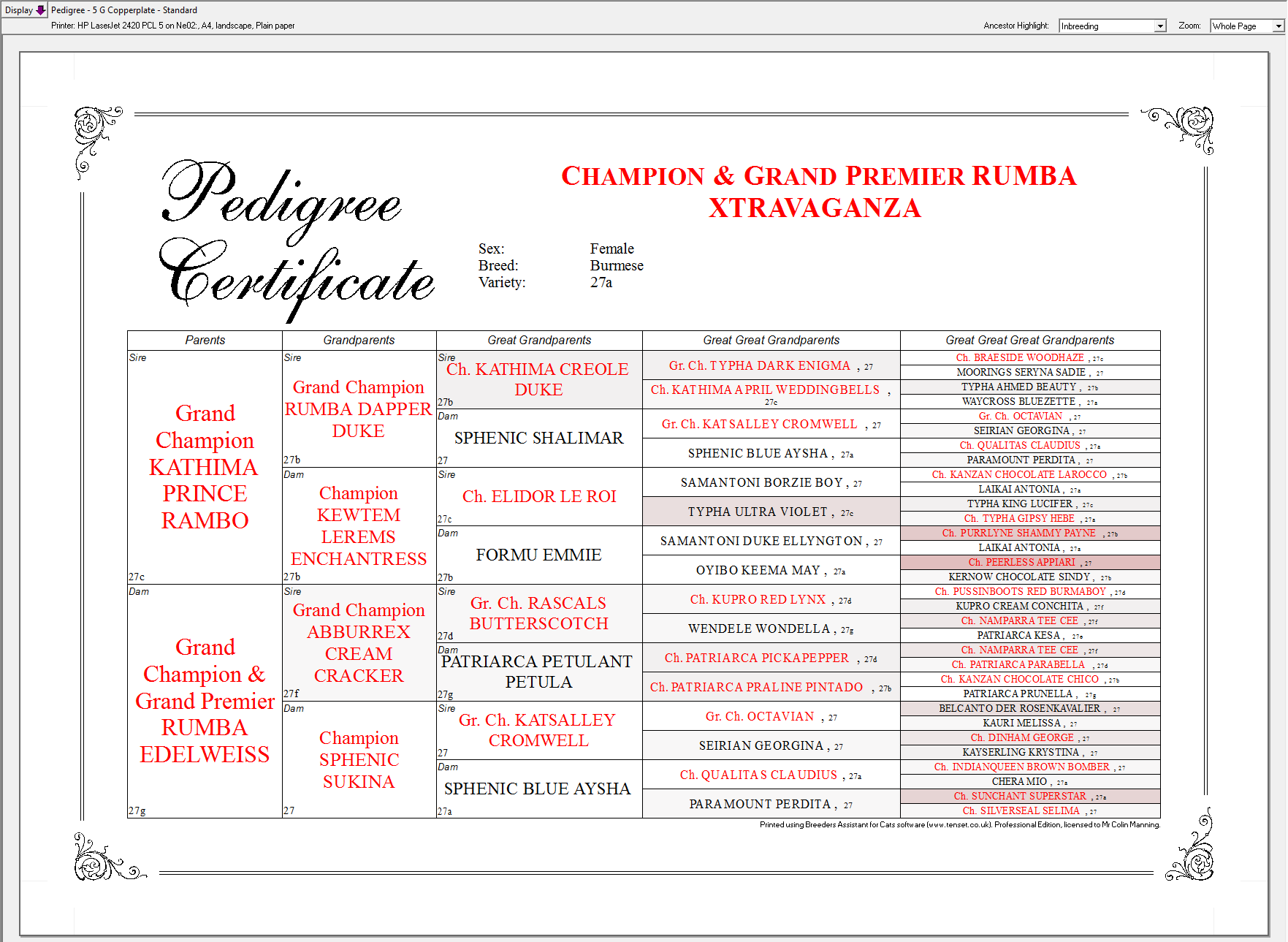
It is also possible to highlight common ancestors (i.e. those ancestors that appear on both the sire and dam sides of the displayed pedigree). Select Common from the Ancestor Highlight pulldown above the pedigree in the main window.
With the Professional and Extended Editions ancestors can be highlighted within a pedigree according to their level of inbreeding. Select any of the Inbreeding options from the Ancestor Highlight pulldown above the pedigree in the main window.
Ancestor Circularity Detection (Self-Parenting)
An ancestory circularity (self-parenting error) occurs when an individual appears within its own ancestors, i.e. you have entered pedigree data where an animal is claimed to be descended - via other ancestors - from itself. Whilst that is impossible in nature, it is possible if you have an error in your pedigree data.
Breeders Assistant has an inbreeding analysis report that will describe the 'loop' in your data. E.g.:
Circularity detected in ancestors at G14-18: Arripay Bigfoot -> Mylady Kazoo -> Ch. Arripay Handsome JW -> Am. & UK Ch. Arripay Sunshine JW -> Arripay Bigfoot
This report is saying that Arripay Bigfoot is descended from itself - and the report shows the other ancestors that make up the 'loop' in the data. It gives the information needed to fix these errors.
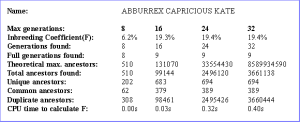
Inbreeding Analysis
An 'inbreeding analysis' report lists the inbreeding for a chosen animal, computed to varying depths of ancestors, and lists associated information such as the depth at which the furthest ancestor was found, how many generations of ancestors were full, the total number of ancestors found, the numbers of unique ancestors and common ancestors (to both sides of the pedigree).
1. Mating certificates are only available with the cat, dog and 'Generic' versions of Breeders Assistant.